Post Processing & Data Analysis
TDTbin2mat and the MATLAB SDK

Exporting data from Synapse into MATLAB is simple with the TDT MATLAB SDK. The main importing function of the MATLAB SDK is TDTbin2mat. The main argument for TDTbin2mat is the full file path to the data block that you want to import. Synapse makes copying this file path easy via the History dialog. See this Lightning Video to see this importing sequence. You can also copy the block file path via Windows Explorer.
The TDT Python Package

Data can also be easily imported into Python 3 using the tdt package. If
you already have Python 3 installed, you can add the tdt package in your
cmd window: pip install tdt
Link to Python Package and SDK:
Release notes and select examples
MATLAB and Python Workbook Examples
TDT aims to help customers as much as possible with easy data import and analysis. We understand that not all customers have extensive MATLAB or Python experience, so we created fully-commented workbook examples that demonstrate how to do basic, but interesting operations with MATLAB or Python code. These examples are not intended to serve as a complete pipeline for your data analysis - please use wisely.
Link to MATLAB Workbook examples
Link to Python Notebook examples
Fiber Photometry Epoch Averaging example (MATLAB)
Fiber Photometry Epoch Averaging example (Python)
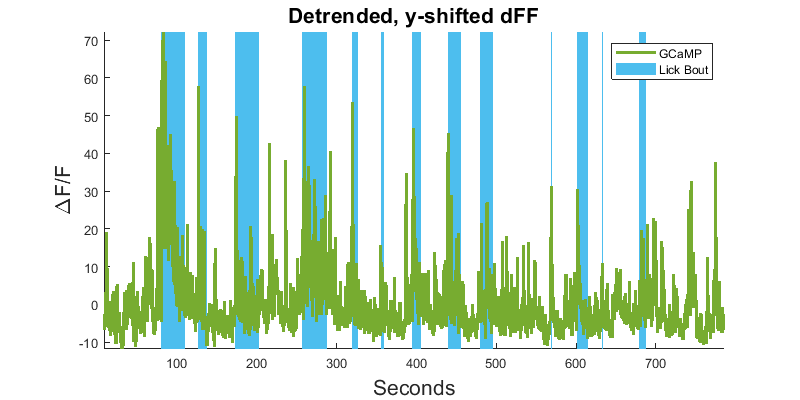
Lick Bout Epoc Filtering (MATLAB)
Lick Bout Epoc Filtering (Python)
If you have other scripting needs, please reach out to TDT Tech Support at support@tdt.com.
View Data in OpenScope
For a first-pass replay of data, you can view any Synapse recording in OpenScope. This also takes advantage of the Synapse History dialog. OpenScope has extra features that make jumping around the data fast and intuitive. You can also use the Video Viewer feature to replay videos with the timestamp of each frame.
If you want to use a Windows computer that does not have Synapse on it, then you can download TTankMin from here and use the password 'killerbee' for intall. This will let you use OpenScope on your PC. If you have trouble locating OpenScope, try searching 'Scope' in Windows.
Note that you must preserve a Tank and Block structure to view data in OpenScope. You may need to add in .ini files for Scope to recognize your tanks and block (only needed if Scope is not seeing your data). The easiest way to transfer a Tank folder with Blocks inside is to zip the Tank folder on the original PC and then unzip on the PC with OpenScope.
Video Review and Scoring
Please visit our guide on Video Review and Scoring for complete information about how to review your recorded videos alongside photometry data.
Exporting to ASCII (CSV) Format
Customers sometimes like to export their data to CSV files for import into Excel, Prism, or other software.
The easiest way to handle this would be to import your data into MATLAB or Python first and then export your individual streams or a matrix of streams as a CSV file. In Python, you can use the read_block export function. In MATLAB you can use the writematrix function. TDT also has an exporting tool called TDTexport. You can read more about the read_block and TDTexport options in our Data Exporting page.
Importing into MATLAB or Python first has the added advantage of allowing you to down sample your data before importing into your CSV reader. Excel, for example, has a row and column limit that often prevents users from importing their full continuous raw data as a single row or column because there are too many sample points.
You can also use OpenBrowser to export to ASCII format.
Note
This video demonstrates exporting to an EDF file format, not ASCII.
More Resources
Here are some common resources that customers may find helpful as they work to understand fiber photometry and conduct experiments.
Select fiber photometry papers:
- Lerner et al. 2015 https://dx.doi.org/10.1016/j.cell.2015.07.014
- Calipari et al. 2016 https://doi.org/10.1073/pnas.1521238113
- Knight et al. 2015 https://dx.doi.org/10.1016/j.cell.2015.01.033
- Barker et al. 2017 https://doi.org/10.1016/j.celrep.2017.10.066
Fiber photometry community forum